

Tobias Madl

Projects within the DK-MCD
| Structural and functional mechanisms in the intersection of p53 and Wnt signaling | Hermann Habacher |
Nuclear receptor signalling in immune-metabolic functions of cerebrovascular endothelial cells |
Magdalena Lang graduated |
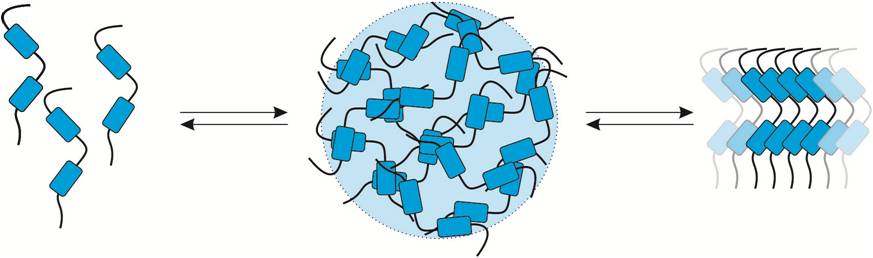
Regulation of nuclear import and phase separation |
Aneta Lenard graduated |
Regulation of nuclear import and phase separation |
Sinem Usluer graduated |
Molecular characterization of TNPO3 cargo recognition and liquid-liquid phase separation (LLPS) behavior of RSY containing splicing factors |
Qishun Zhou graduated |
Research interest
Our group investigates the molecular mechanisms how disordered proteins mediate signal transfer via an intricate network of protein interactions and post-translational modifications. We work at the interface of structural biology, biophysics, cell biology, and medicine to study the molecular mechanisms linking key regulatory processes involved in signaling and metabolism. Nuclear magnetic resonance (NMR) and biophysical methods are the primary tools employed in the laboratory to study the structural and dynamical properties of signaling proteins, metabolites and their interactions under native-like solution conditions. The efforts focus primarily on the highly conserved Wnt signaling pathway and intersected pathways, forkhead transcription factors, and the biogenesis of RNA-binding proteins. Distortions in these pathways lead to a plethora of diseases, including cancers, neurological and metabolic disorders, and are linked to aging. The group aims to reveal the molecular mechanisms underlying interactions of disordered proteins to provide insight into the intricate link between their function, regulation, and human diseases. We use NMR-based metabolic phenotyping to obtain systemic insight into the molecular outcome of distortions in signaling pathways in cell-based assays, in vivo model systems, and patients. This allows to bridge the gap between atom and patient and vice versa and to obtain new insights into disease mechanisms and diagnosis.
To be able to study large and dynamic biomolecular complexes involved in signal transduction, the team employs and develops an integrated structure determination protocol in which they combine NMR spectroscopy, SAXS/SANS, complementary techniques (i.e. MS, EM, FRET), and modeling strategies especially suited to study the structure of large and dynamic biomolecules and biomolecular complexes. Next to molecular dynamics programs widely used in the NMR community, our computational focus is on the integration of novel techniques in the Rosetta de novo structure prediction framework. Several approaches were developed and applied to challenging biological systems in the last few years. |
|
Curriculum vitae
| 1998 - 2004 | Studies of Chemistry, Physics, History and Computational Sciences at the University Graz, Austria | |
| 1999 - 2000 | Laboratory assistant, Landeskrankenhaus Graz (hospital), Austria | |
| 2002, 2004 | First Diplomas within the enrolled programs (comparable to Bachelor of Science) | |
| 2003 - 2004 | Diploma thesis, Institute of Chemistry, University of Graz, Austria | |
| 2004 - 2007 | PhD thesis, University of Graz, Austria (DOC Fellowship of the ÖAW) | |
| 2007 | Postdoc in Structural Biology at the Institute of Chemistry, University of Graz, Austria | |
| 2007 - 2010 | Postdoc in integrative structural biology of protein-RNA complexes at the Department of Chemistry of the Technical University and at the Institute of Structural Biology at the Helmholtz Center Munich, Germany (Schrödinger Fellowship of the FWF, EMBO-LTF) | |
| 2010 - 2011 | PostDoc, Utrecht University, The Netherlands | |
| 2012 | APART fellow at Utrecht University, The Netherlands and University Graz, Austria | |
| 2012 - 2016 | Emmy Noether and BioSysNet research group leader, TUM Junior Fellow, Member of the CIPSM excellence cluster, Member of the Junior Scientist Program (Tenure Track), at Technical University and Helmholtz Zentrum Munich (HMGU), Germany | |
| 2015 - | Visiting Professor, Fujian Institute of Research on the Structure of Matter (FJIRSM), Chinese Academy of Sciences, Xiamen, China | |
| 2015 - 2023 | Associate Professor at the Institute of Molecular Biology and Biochemistry, Medical University Graz | |
| 2016 | Habilitation in Molecular Biology | |
| 2016 - | Full Member of BioTechMed-Graz | |
| 12/2023 - | Head of the Division of Medicinal Chemistry | |